I have put together an Alfred workflow – this one searches the HGNC database for genes! I have converted the text database from the HGNC website and configured it for full text search using sqlite. This allows you to lookup genes by their UCSC, Entrez, Vega, Ensembl, and many other identifiers very quickly.
Download the latest release
Usage
Full text search of the HGNC database

Information and links are provided for individual genes
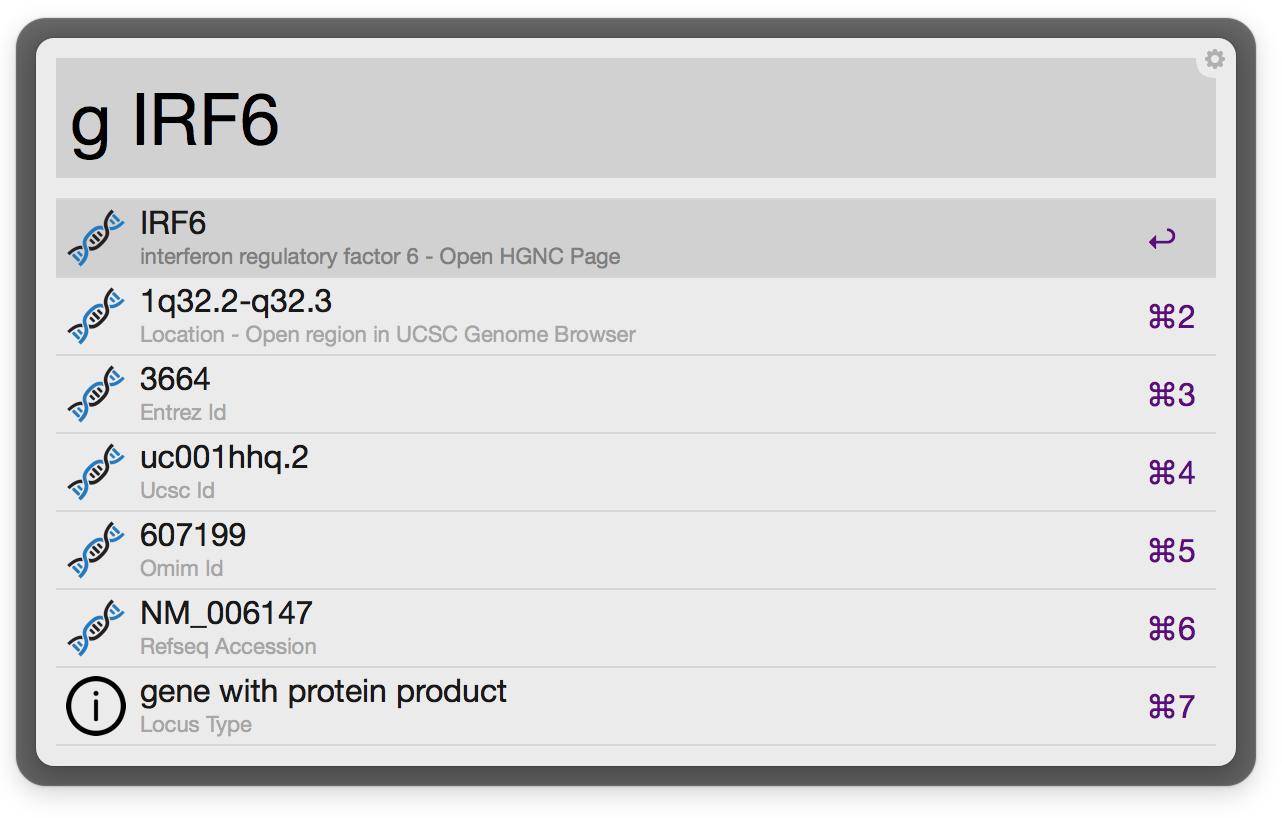
Feedback
Please provide feedback. Specifically:
- What other gene IDs should be displayed by default? (You can currently search for any)
- What other sites would you like to be able to navigate to.
- Is there additional information that should be folded in that would be useful?